Related Publications
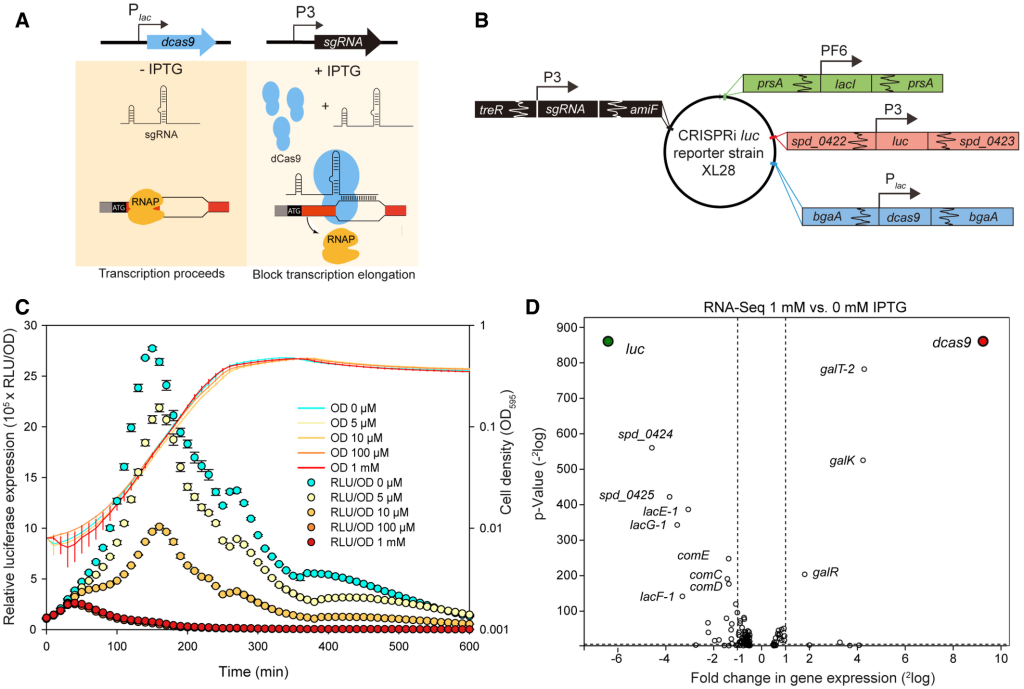
- Liu, X., Gallay, C., Kjos, M., Domenech, A., Slager, J., Kessel, S.P. van, Knoops, K., Sorg, R.A., Zhang, J.-R., and Veening, J.-W. (2017). High‐throughput CRISPRi phenotyping identifies new essential genes in Streptococcus pneumoniae. Molecular Systems Biology 13, 931. https://doi.org/10.15252/msb.20167449. (cover paper) (CRISPRi phenotyping)
- de Bakker, V., Liu, X., Bravo, A.M., and Veening, J.-W. (2022). CRISPRi-seq for genome-wide fitness quantification in bacteria. Nat Protoc 17, 252–281. https://doi.org/10.1038/s41596-021-00639-6. (CRISPRi-seq)
- Dénéréaz, J., Eray, E., Jana, B., De Bakker, V., Todor, H., Van Opijnen, T., Liu, X., and Veening, J.-W. (2024). Dual CRISPRi-Seq for genome-wide genetic interaction studies identifies key genes involved in the pneumococcal cell cycle. Preprint, https://doi.org/10.1101/2024.08.14.607739. (Dual CRISPRi-seq)
- Jana, B., Liu, X., Dénéréaz, J., Park, H., Leshchiner, D., Liu, B., Gallay, C., Zhu, J., Veening, J.-W., and van Opijnen, T. (2024). CRISPRi-TnSeq maps genome-wide interactions between essential and non-essential genes in bacteria. Nat Microbiol 9, 2395–2409. https://doi.org/10.1038/s41564-024-01759-x. (CRISPRi-Tn-seq)
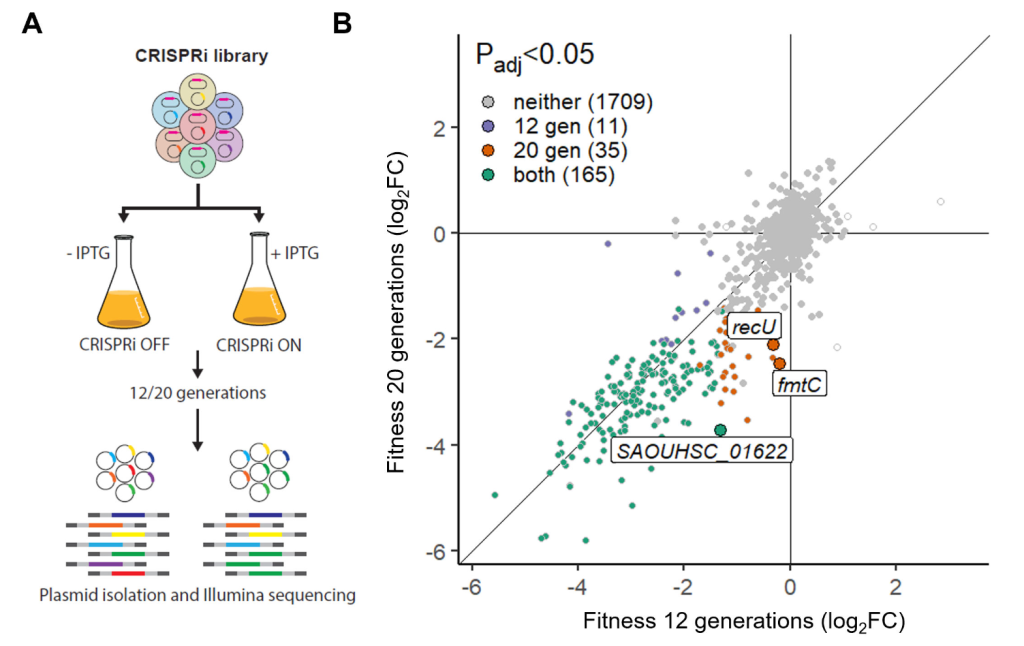
- Zhou, Y., Song, Y., Zhang, Y., Liu, X., Liu, L., Bao, Y., Wang, J., and Yang, L. (2024). Azalomycin F4a targets peptidoglycan synthesis of Gram-positive bacteria revealed by high-throughput CRISPRi-seq analysis. Microbiol Res 280, 127584. https://doi.org/10.1016/j.micres.2023.127584. (CRISPRi-seq)